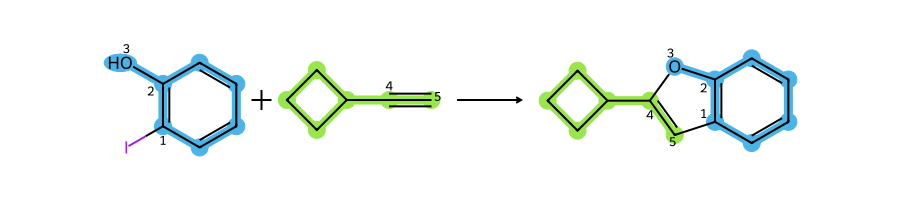
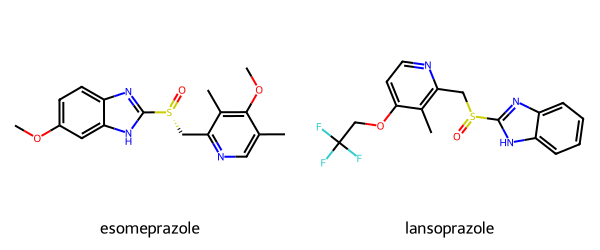
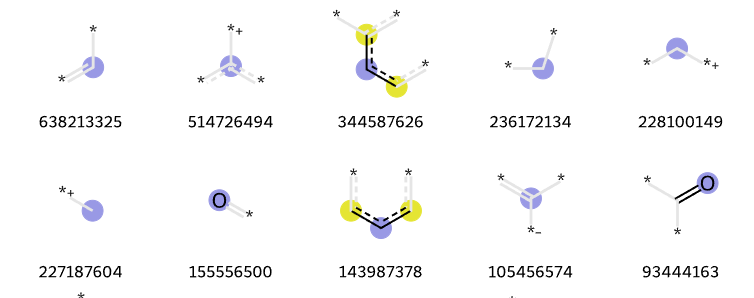
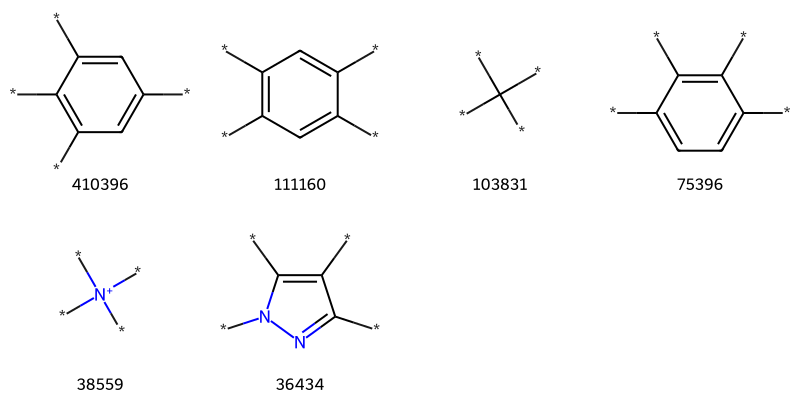
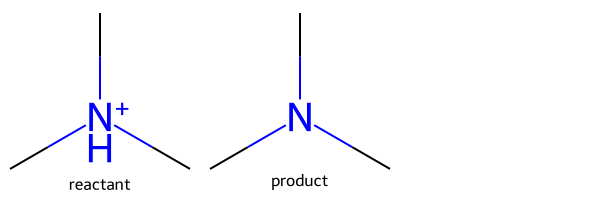
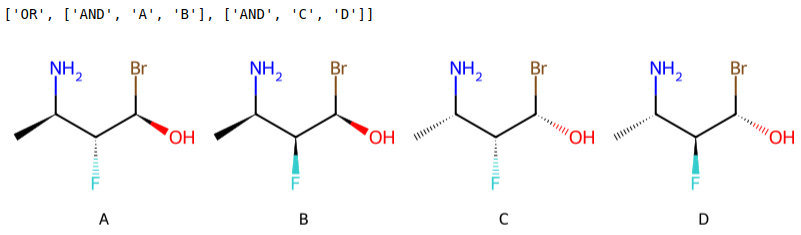
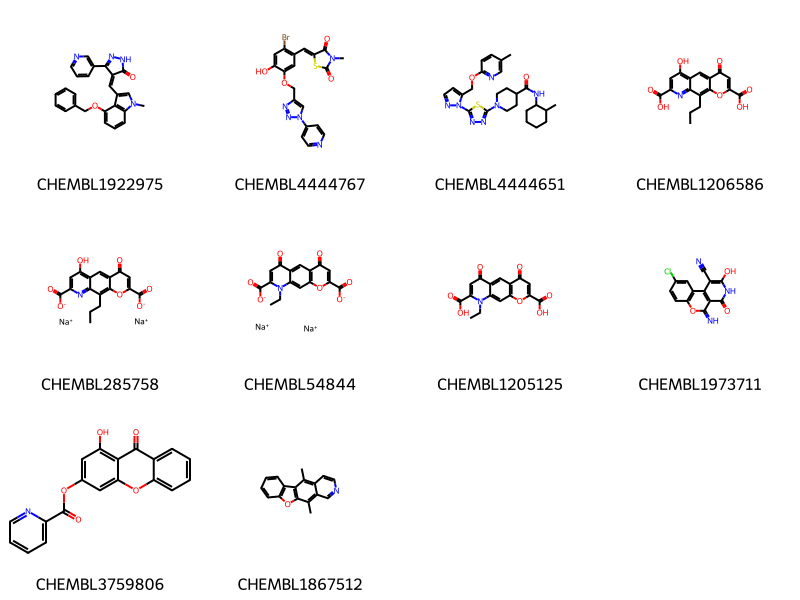
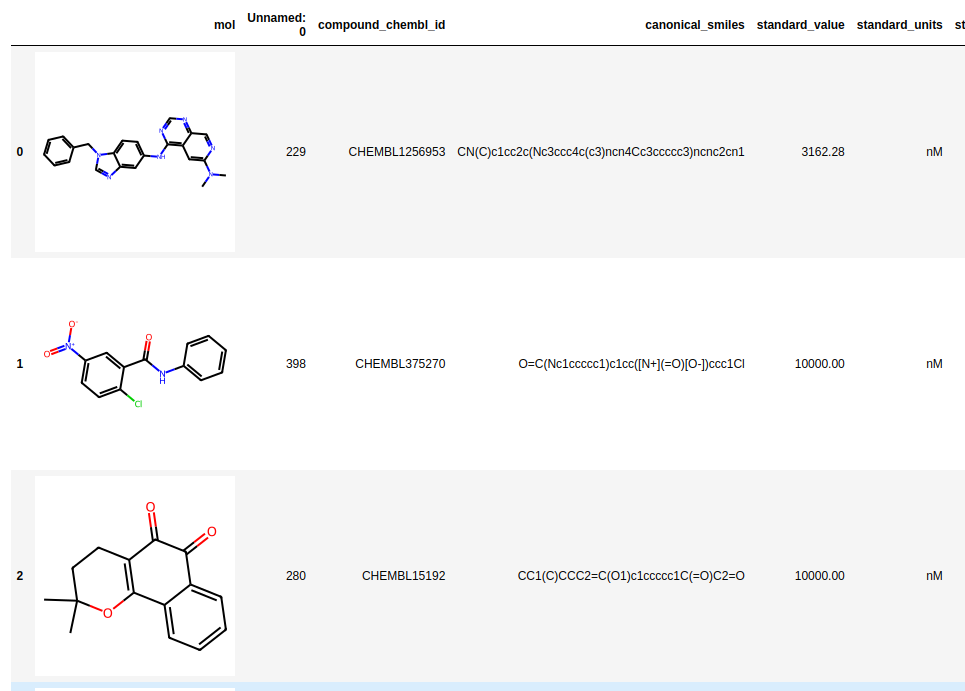
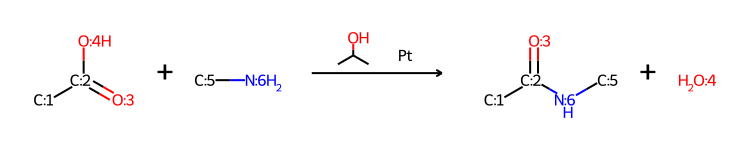
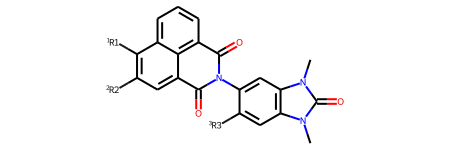
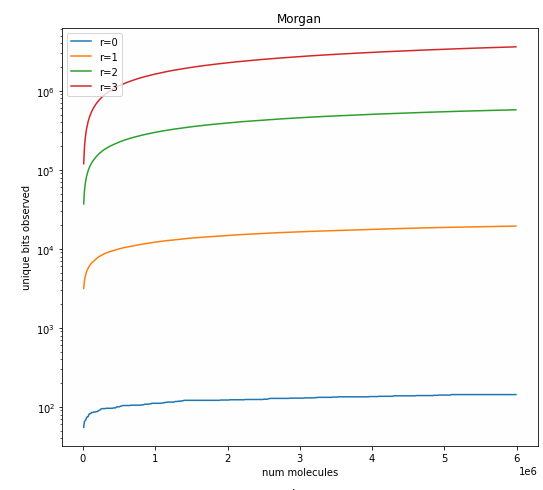
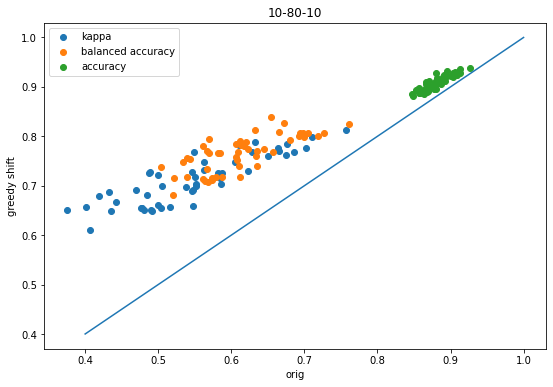
Creating a patent data set
datasets
exploration
patents
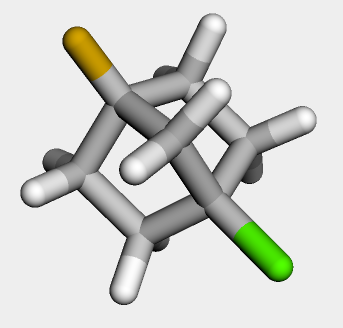
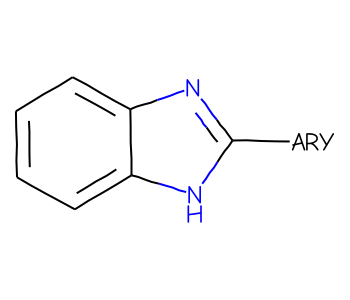
About tetrahedral chirality in the RDKit
tutorial
documentation
stereochemistry
Building synthon spaces with combinatorial reactions
tutorial
documentation
Working with the LOBSTER Data set II
datasets
3d
similarity
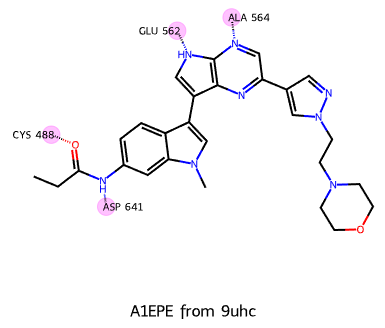
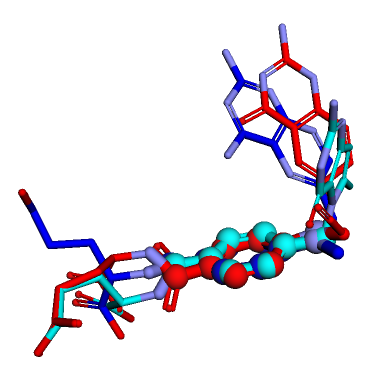
Drawing simple protein–ligand interaction diagrams with the RDKit
drawing
exploration
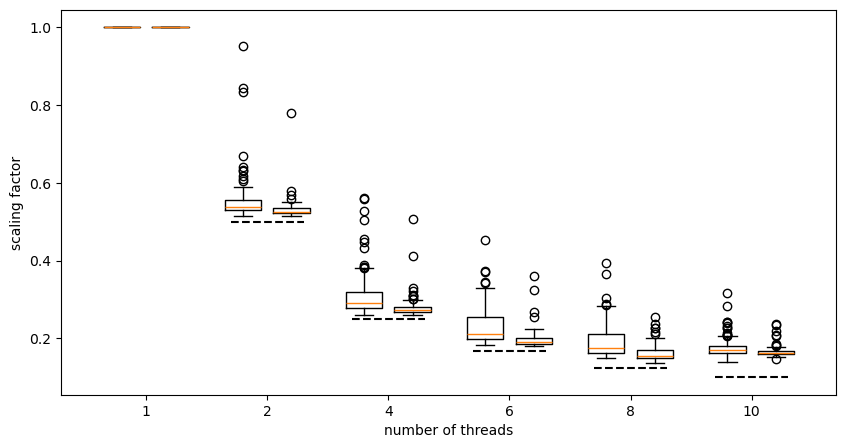
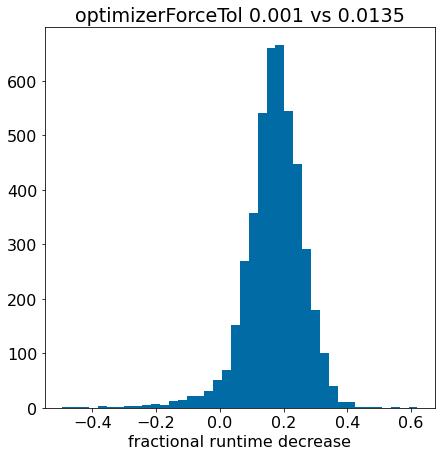
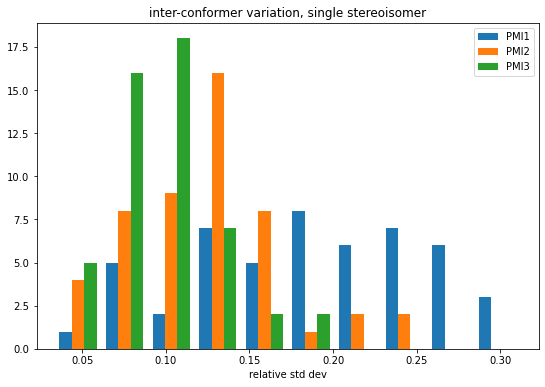
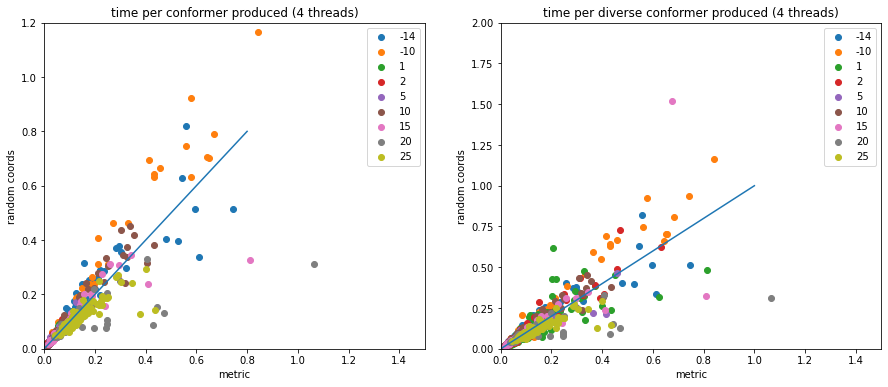
Scaling conformer generation
conformers
questions
technical
Controlling the RDKit’s logging behavior
documentation
technical
An LLM experiment
exploratoration
LLMs
Please stop saying “The Tanimoto similarity is”
similarity
fingerprints
rants
Picking a small set of diverse molecules from a large set
similarity
fingerprints
tutorial
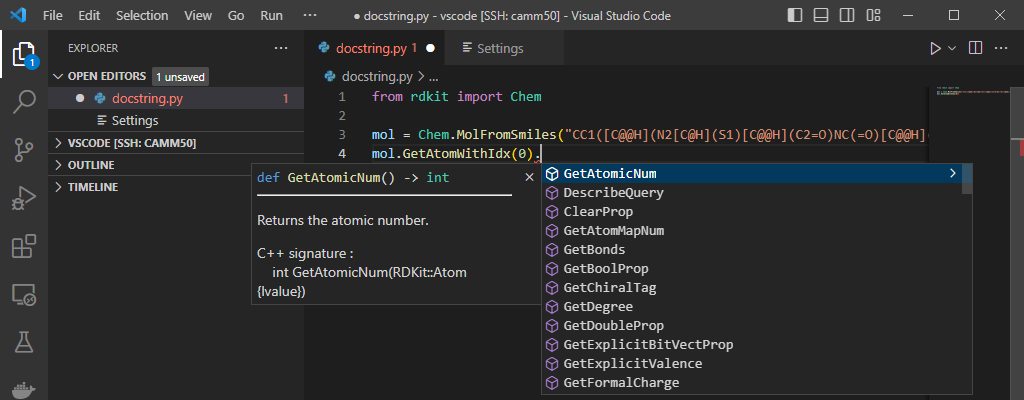
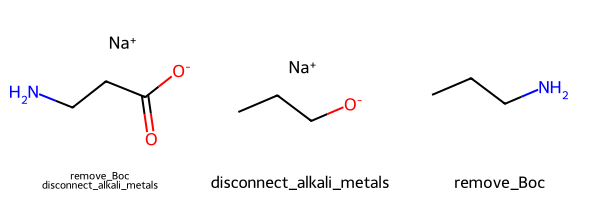
Sanitization options and molecule parsing.
documentation
technical
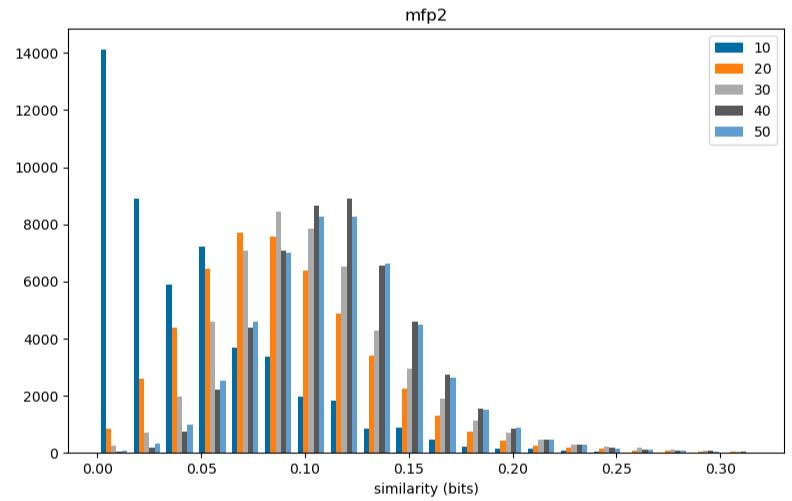
The impact of molecular size on similarity.
reference
similarity
fingerprints
Murcko Scaffolds Tutorial
tutorial
guest post
Using custom fingerprints in PostgreSQL
cartridge
fingerprints
tutorial
Explaining enhanced stereochemistry
prototypes
tutorial
stereochemistry
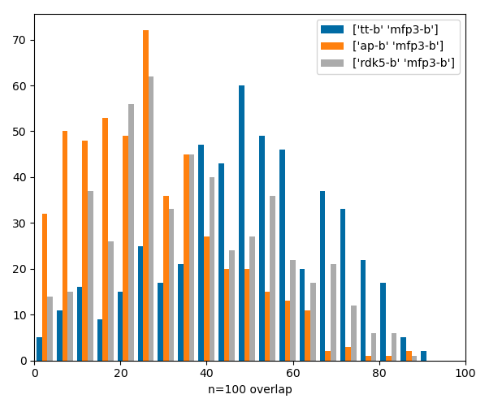
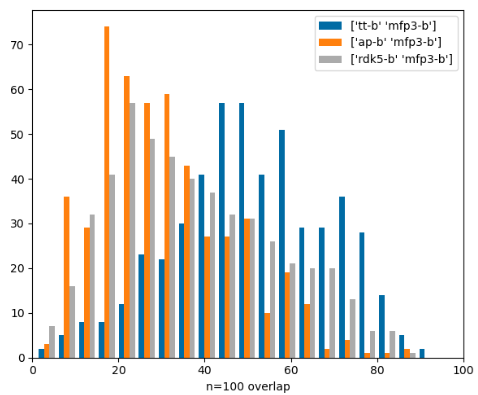
Similarity-search hitlist overlap part 2
reference
similarity
fingerprints
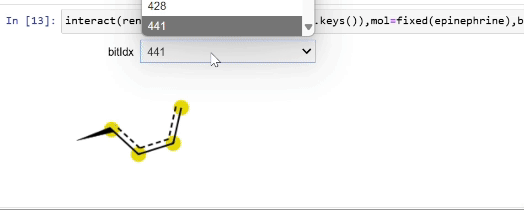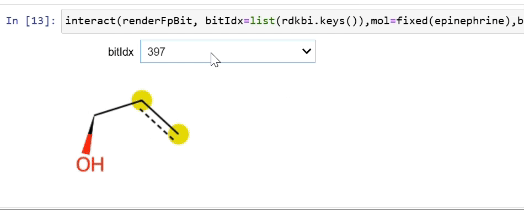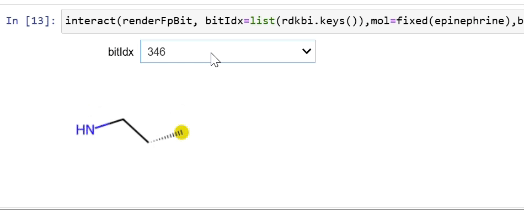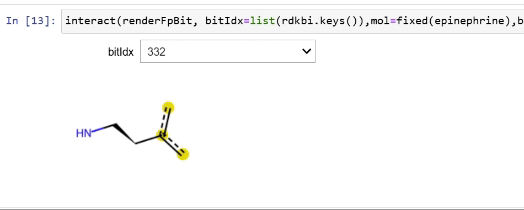
Rendering fingerprint bits
tutorial
drawing
fingerprints
Similarity-search hitlist overlap
reference
similarity
fingerprints
Building a similarity comparison set
reference
similarity
Tuning substructure queries
tutorial
substructure
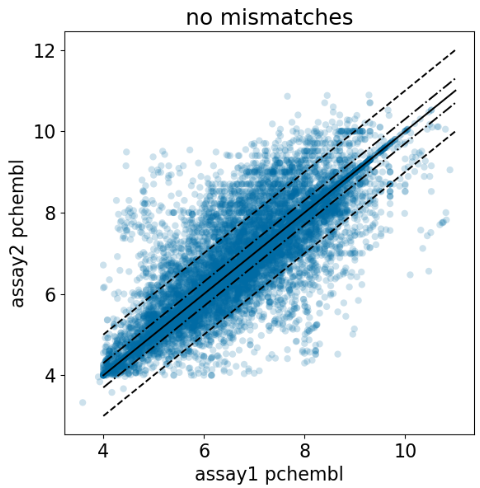
ChEMBL Document Similarity
exploration
similarity
Introducing Synthon Searching
release
tutorial
guest post
Using the ACS1996 drawing style in PandasTools
documentation
tutorial
Additive fingerprints
exploration
similarity
fingerprints
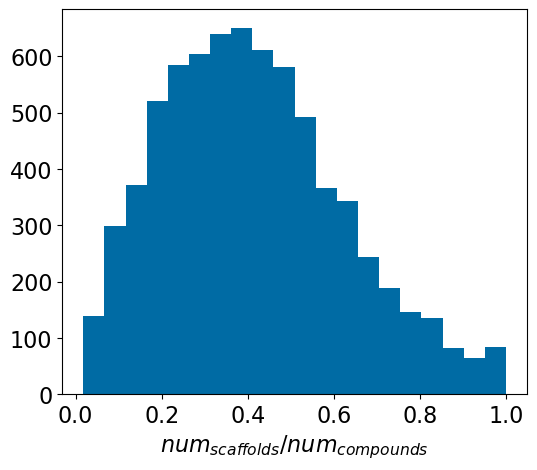
The problem(s) with scaffold splits, part 1
machine learning
rants
reference
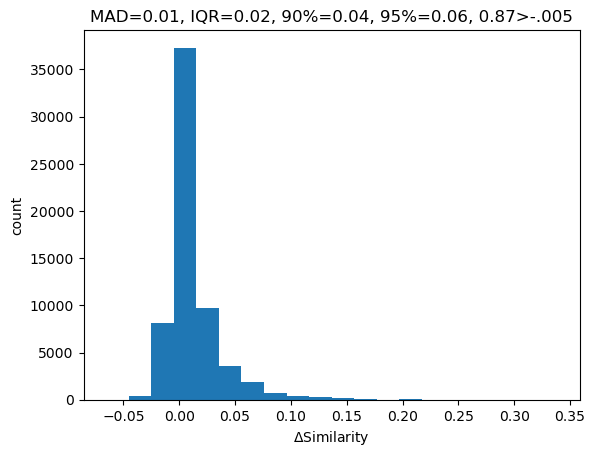
The impact of single-atom changes on similarity
fingerprints
reference
exploration
Calculating the SA_Score and NP_Score descriptors
tutorial
documentation
descriptors
Intro to Stereo Groups and Enhanced Stereochemistry
tutorial
stereochemistry
documentation
Plotting rows and columns of molecules with MolsMatrixToGridImage
release
tutorial
guest post
Variability of x-fold cross validation results
machine learning
exploration
Binary molecules and the cartridge
cartridge
tutorial
exploration
Colliding bits III, expanded
reference
fingerprints
Colliding bits II, revisited
reference
fingerprints
Timing methods for serializing molecules
reference
optimization
Dealing with multiconformer SD files
3d
conformers
tutorial
Optimizing conformer generation parameters
3d
conformers
optimization
A Ternary GHOST
exploratory
machinelearning
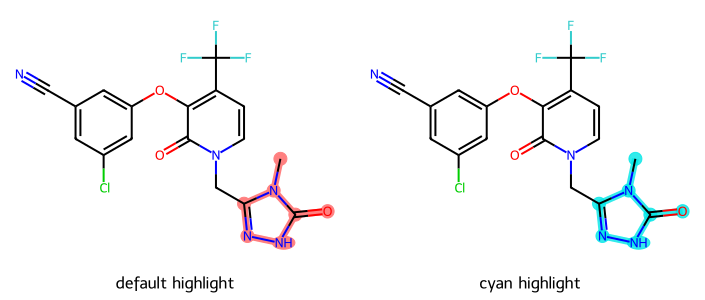
R-Group Decomposition and Highlighting
tutorial
prototypes
drawing
rgd
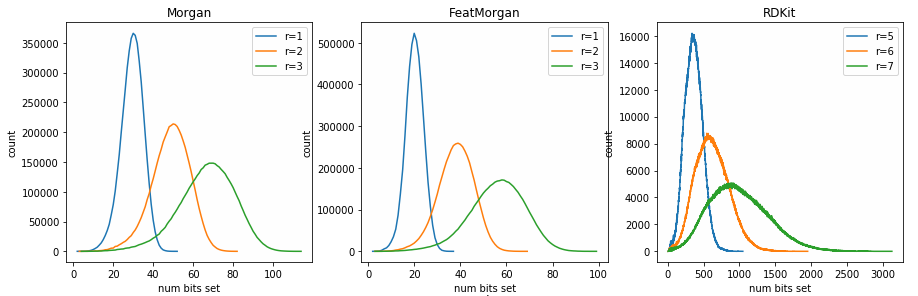
Looking at the number of bits set by different fingerprints
fingerprints
reference
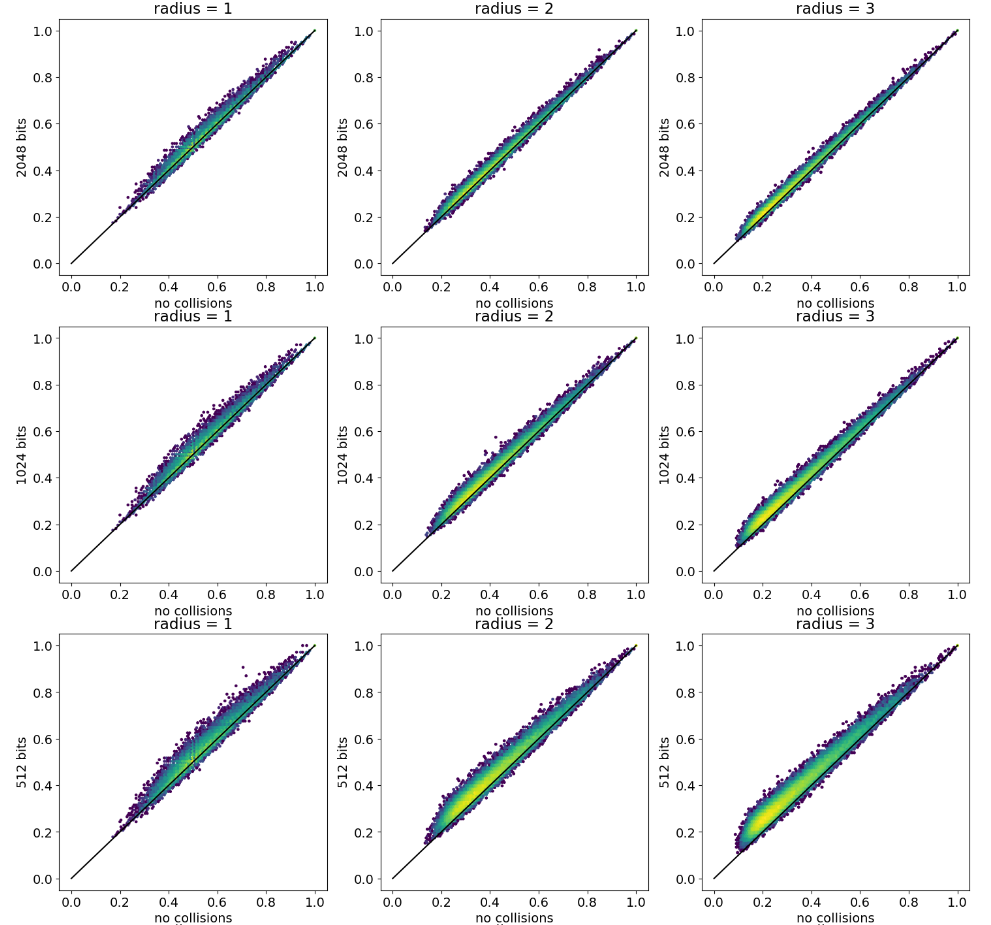
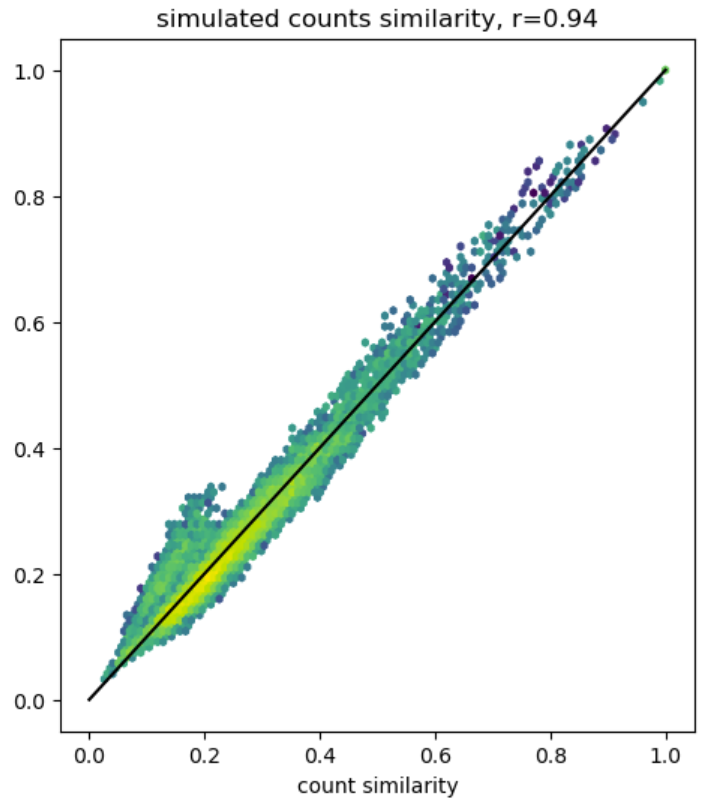
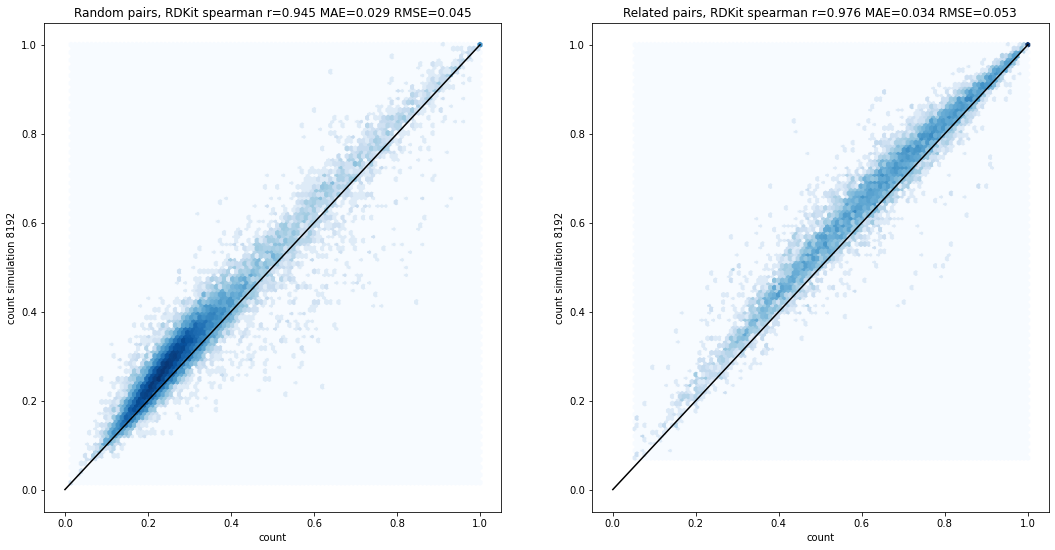
Simulating count fingerprints
fingerprints
technical
reference
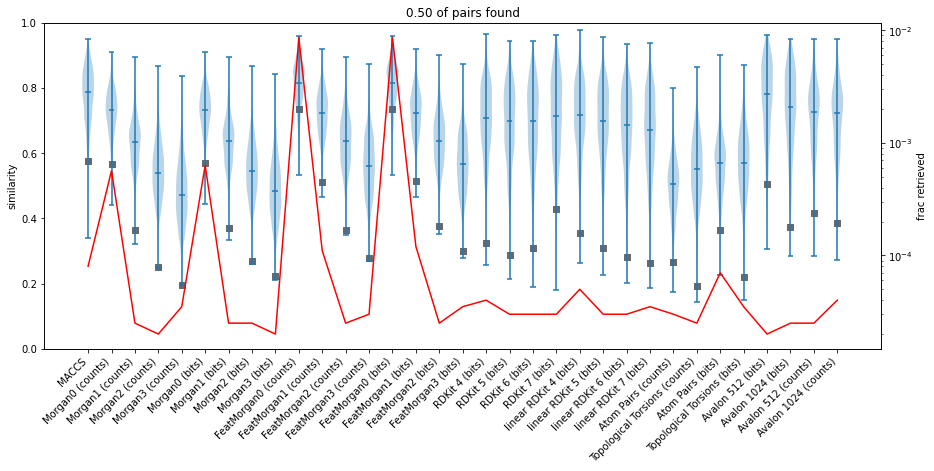
Fingerprint similarity thresholds for database searches
similarity
reference
Thresholds for “random” in fingerprints the RDKit supports
fingerprints
similarity
reference
Looking at random-coordinate embedding
conformers
exploration
3d
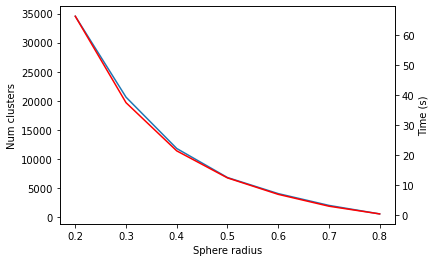
Sphere exclusion clustering with the RDKit
similarity
tutorial
guest post
No matching items